New biomolecules to detect bacteria
The ICC is developing cost-effective detection molecules for future rapid diagnostic testing methods.
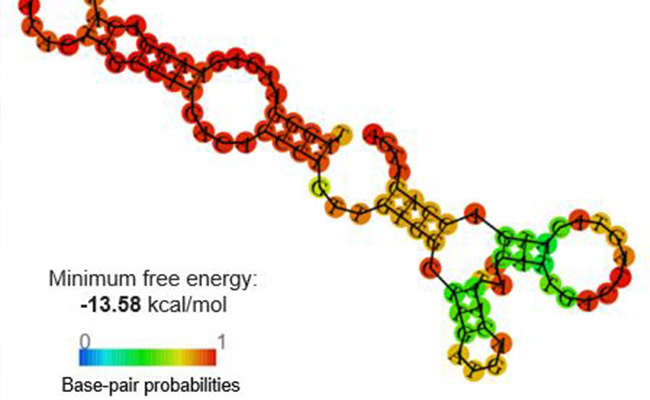
Secondary structure of the developed DNA aptamer for Enterococcus faecalis
The ICC Water & Health is developing cost-effective detection molecules for bacteria - so-called DNA aptamers - which offer great potential for future rapid diagnostic testing methods.
Rapid cell detection
Rapid direct cell detection methods, such as (online-) flow cytometry or microscopy, can currently only be used to a limited extent for water quality testing. The reason behind is that they cannot differentiate between harmless and health-relevant bacteria. Inexpensive recognition molecules, such as aptamers, coupled with fluorescent dyes could provide a remedy and serve as tools for targeted cell labeling in the future.
Claudia Kolm has now generated DNA aptamers for fecal bacteria in water and established the aptamer development technology at TU Wien in the research group Environmental Microbiology and Molecular Diagnostics of Andreas Farnleitner's assigned working group of Georg Reischer. The method was recently published in the scientific journal "Scientific Reports".
Customized recognition molecules from DNA building blocks
Aptamers are short, synthetically produced DNA or RNA molecules which, due to their three-dimensional structure, recognize and specifically bind target molecules. Similar to antibodies, binding takes place according to the lock-and-key model. In this way aptamers bind to specific proteins and complex cell surface structures. However, specific aptamers can also be generated for small molecules, such as antibiotics or various toxins.
Research and industry are increasingly recognizing the potential of aptamers as these "chemical antibodies" are more robust and much easier to produce, without the risk of batch-to-batch variations. Unlike antibodies, aptamer development is also not dependent on animals and cell lines. They are generated in vitro and can be adapted for diagnostic end uses. For instance, the conditions under which the aptamer binds its target (buffer, temperature, etc.) can be defined and tuned during the development process.
Evolution in a test tube
The starting point of aptamer development and the published method with Enterococcus faecalis cells as target was a combinatorial DNA library with ~1015 different single-stranded DNA molecules. To screen this huge DNA pool for suitable candidates, a process similar to natural evolution is used. DNA molecules that bind to the target bacterium are selected and enriched. Over several rounds with increasing selection pressure, good binders are separated from non- and weak binders. In combination with modern sequencing methods and developed bioinformatic data analysis tools, it was possible to enrich and identify a specific aptamer for Enterococcus faecalis in the DNA pool. Tests with closely related and other bacterial species found in fecal polluted waters showed no cross-reactivity.
Both the generated aptamer and the established aptamer development platform offer great potential for further research & development. In a project funded by the state of Lower Austria (Life Science Call LS17-007) the development platform is now just to generated DNA aptamers for Vibrio cholerae - the causative agent of cholera.
Original publication:
Kolm C., Cervenka I., Aschl U.J. et al. DNA aptamers against bacterial cells can be efficiently selected by a SELEX process using state-of-the art qPCR and ultra-deep sequencing. Sci Rep 10, 20917 (2020)
Funding source:
- Austrian Academy of Sciences (DOC Fellowship, Claudia Kolm)
- Niederösterreichische Forschungs- und Bildungsges.m.b.H.
Contact details:
Dr. Georg Reischer | ICC Water&Health | TU Wien – Institute of Chemical, Environmental & Bioscience Engineering | georg.reischer@tuwien.ac.at
Dr. Claudia Kolm | ICC Water&Health | TU Wien – Institute of Chemical, Environmental & Bioscience Engineering | claudia.kolm@tuwien.ac.at
